Page 45 - CSIR-IGIB Annual Report 2020-21
P. 45
Ribosomal methylation and antimicrobial function of decreasing pH identified a highly
resistance stable conformation of Rv2966c at acidic pH (pH
Owing to the central role of protein synthesis in 4.0) providing clues to the structure and stability
cellular function, ribosomes are the most of Rv2966c, at low pH in the cell. We also
common targets of antibiotics. The ribosome determined the crystal structure of RsmG-
harbors several modifications viz., homolog (Rv3919c) in complex with sinefungin
pseudouridylation, ribose methylation and base that helped identify the binding site. A deletion
methylation in both 16S as well as 23S rRNA. mutant of the RsmG-homolog was generated in
These methylations are brought about by highly M. smegmatis mc 155 that confirmed its role in
2
site-specific methyltransferases. Any alteration increased resistance to ribosome-targeting
in the methylation patterns affects ribosome antibiotics, specifically to streptomycin.
function and alters the response to drugs in
bacteria. However, the exact mechanism of the
highly specific mode of substrate binding and
recognition is unclear. In this work, we
investigate the specific modes of recognition by
16S rRNA methyltransferases of M. tuberculosis
and their roles in modulating drug response due
to alterations of methylation patterns.
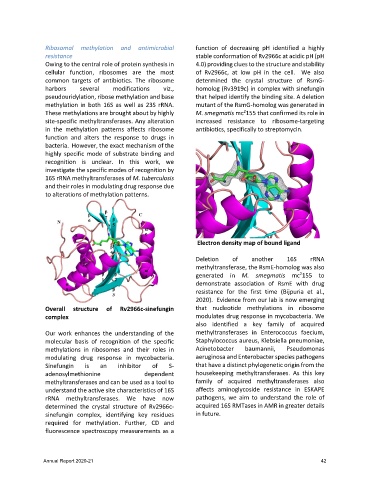
Electron density map of bound ligand
Deletion of another 16S rRNA
methyltransferase, the RsmE-homolog was also
generated in M. smegmatis mc 155 to
2
demonstrate association of RsmE with drug
resistance for the first time (Bijpuria et al.,
2020). Evidence from our lab is now emerging
Overall structure of Rv2966c-sinefungin that nucleotide methylations in ribosome
complex modulates drug response in mycobacteria. We
also identified a key family of acquired
Our work enhances the understanding of the methyltransferases in Enterococcus faecium,
molecular basis of recognition of the specific Staphylococcus aureus, Klebsiella pneumoniae,
methylations in ribosomes and their roles in Acinetobacter baumannii, Pseudomonas
modulating drug response in mycobacteria. aeruginosa and Enterobacter species pathogens
Sinefungin is an inhibitor of S- that have a distinct phylogenetic origin from the
adenosylmethionine dependent housekeeping methyltransferases. As this key
methyltransferases and can be used as a tool to family of acquired methyltransferases also
understand the active site characteristics of 16S affects aminoglycoside resistance in ESKAPE
rRNA methyltransferases. We have now pathogens, we aim to understand the role of
determined the crystal structure of Rv2966c- acquired 16S RMTases in AMR in greater details
sinefungin complex, identifying key residues in future.
required for methylation. Further, CD and
fluorescence spectroscopy measurements as a
Annual Report 2020-21 42